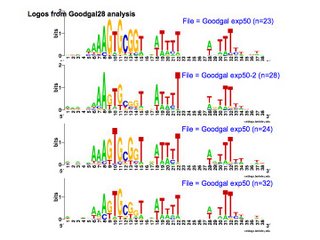
 First, I can use the Gibbs motif sampler to search it for USS-like patterns. This provides one direct estimation of the bias of the uptake machinery. Here are some replicate results. All of these searches used the same dataset, but random differences in the search runs produced different datasets, which produced different logos. I haven't gone back and compared the outputs to see how many of the same sequences they found, but that will only take a few minutes to do, using Unix's 'diff' command.
First, I can use the Gibbs motif sampler to search it for USS-like patterns. This provides one direct estimation of the bias of the uptake machinery. Here are some replicate results. All of these searches used the same dataset, but random differences in the search runs produced different datasets, which produced different logos. I haven't gone back and compared the outputs to see how many of the same sequences they found, but that will only take a few minutes to do, using Unix's 'diff' command.Second, I can find out how well the uptake of these sequences is predicted by their degree of match to the genomic USS pattern. I know that I can use a program called PatSer (for Pattern Search, I guess) on the RSA Tools site to do this. It constructs a scoring matrix and then scores sequences you give it. The matrix will be constructed from the probability matrix that the Gibbs searches produce, but I need to ask one of the graduate students to help me do this. Once I have the scores, I'll plot a graph of molecules taken up as a function of PatSer score - a strong correlation would support the hypothesis that biased uptake is responsible for the accumulation of USS in the genome.
















runescape money runescape gold runescape money runescape gold wow power leveling wow powerleveling Warcraft Power Leveling Warcraft PowerLeveling buy runescape gold buy runescape money runescape items runescape gold runescape accounts runescape gp dofus kamas buy dofus kamas Guild Wars Gold buy Guild Wars Gold runescape accounts buy runescape accounts runescape lotro gold buy lotro gold lotro gold buy lotro gold lotro gold buy lotro gold lotro gold buy lotro gold runescape money runescape power leveling runescape money runescape gold dofus kamas cheap runescape money cheap runescape gold Hellgate Palladium Hellgate London Palladium Hellgate money Tabula Rasa gold tabula rasa money lotro gold
ReplyDeletebuy lotro gold Tabula Rasa Credit
Tabula Rasa Credits