before | uptake
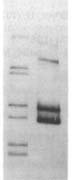 Usually the 'before' pool used in these gels contains end-labeled restriction fragments of a plasmid containing a H. influenzae DNA insert. All the bands are of equal intensity because there are the same number of molecules of each piece of the plasmid. The left lane of the gel figure shows such a 'before' sample. The labeled DNA is then mixed with competent cells, and after a few minutes the DNA that hasn't been taken up is washed away and/or degraded with DNaseI, and DNA is purified from the cells and run in a gel with the 'before' sample. (This 'uptake' DNA prep includes the cells' own DNA, but that's not radioactive so it doesn't show up on the gel.)
Usually the 'before' pool used in these gels contains end-labeled restriction fragments of a plasmid containing a H. influenzae DNA insert. All the bands are of equal intensity because there are the same number of molecules of each piece of the plasmid. The left lane of the gel figure shows such a 'before' sample. The labeled DNA is then mixed with competent cells, and after a few minutes the DNA that hasn't been taken up is washed away and/or degraded with DNaseI, and DNA is purified from the cells and run in a gel with the 'before' sample. (This 'uptake' DNA prep includes the cells' own DNA, but that's not radioactive so it doesn't show up on the gel.)There are two kinds of differences between the before and after lanes. First, some of the 'before' bands are missing (or faint) in the 'uptake' lane. That's because these fragments lack USSs and so were taken up poorly or not at all. Second, faint new bands appear that weren't in the Before sample. These are due to the ligase puzzle.
The authors of the paper said that the new bands appeared because of a ligase enzyme that joins the ends of DNA fragments while they are in the periplasm (the space between the outer and inner membranes. Similar bands were seen in similar experiments in later papers from this lab. A ligase in the periplasm has also been invoked to explain the presence of joined-together DNA fragments recombined into chromosomal DNA.
But the whole idea of a periplasmic ligase seemed a bit odd, as what would a ligase do in the periplasm? There isn't normally any DNA there, and even during DNA uptake there's no obvious role for a ligase.
However, when we did a microarray analysis to identify the genes turned on when cells become competent (see link to CRE-2005 paper in sidebar), we found that one of the genes turned on encodes a ligase with a signal sequence that should target it to the periplasm. Unbeknownst to us, the enzyme had already been well-characterized by biochemists - it's a real ligase, but the ATP-dependent kind typical of some phages, rather than the NAD-dependent kind that acts in DNA replication and repair.
So not only were the early researchers correct in invoking a ligase in the periplasm, but this ligase is specifically turned on when cells are preparing to take up DNA. Consistent with such a role, VanWagoner knocked out the ligase gene and found that transformation was reduced about six-fold. However an undergraduate student in our lab spent last year trying unsuccessfully to see evidence of the ligase activity, and she could not replicate this six-fold reduction.
This ligase needs ATP as a source of energy for its ligation reaction. But as far as we can find out, there is no ATP in the periplasm. In fact, the periplasm contains phosphatases that would cut the 'P' off of any ATP that found its way into the periplasm. One solution would be to have the ligase arrive in the periplasm already loaded with ATP. This is consistent with how such enzymes act - they first form a covalent bond with ATP, and then look for DNA to act on. But I don't know if the machinery that transports enzymes into the periplasm could use a ligase that had already assembled with its ATP. Furthermore, such an 'enzyme' could only act once, and it's hard to imagine that taking up DNA is so important that each molecule is worth sacrificing a whole ligase for.
Bottom line: we still have no idea what role this ligase might play in DNA uptake. If the ligase was essential for DNA uptake, explaining what it accomplishes might be seen as a test of any proposed mechanism of DNA uptake. It's easier to think of roles in the cytoplasm, but all the evidence points to action in the periplasm.
















Wow, that is really interesting. What would a ligase be needed for in DNA uptake? Were all of the experiments done in KW20 strains? I wonder if all strains require the ligase for efficient transformation....
ReplyDeleteI haven't really thought about this before and you've made it sound really interesting. The only way I can imagine a ligase being needed is if uptake of the DNA from the external environment and transport of DNA into the cytoplasm is coupled and part of one continuous process. Then, a break in the DNA in the periplasm could disrupt DNA uptake from outside - I am imagining threading a bit of string through some double glazing - if the strong snaps in the double glazing cavity, it's a bummer, unless you have some little machine in the cavity that can mend the string. I have a vague idea that models of DNA uptake propose that the driving force to pull the DNA into the periplasm lies within the outer membrane, and that would work against an idea of some sort of pulling power coming from proteins situated in the inner membrane. However, I don't think that would preclude some kind of intimate interaction between OM and IM proteins that might require an intact DNA strand suspended across the two for maximum efficiency?
ReplyDeleteMy only other thought is how did authors know those bands always came from assembly of two small bands into one big band, and not digestion of one big band into some smaller bands? I suspect a cursory glance at just one paper will answer that question! Another job for Thursday...
Why did I write that without really sitting and thinking first? It sounds silly now. I guess another possible benefit of a ligase would simply be in providing a long, continuous DNA fragment for the IM machinery to pull through, rather than it having to reinitiate uptake of many, short fragments, which might be more hit and miss. This is really bringing home to me that I can't yet picture the uptake machinery...
ReplyDeleteArghh, here I go again and then I am going to bed. I have Blogorreah. If it costs more ATPs to initiate uptake of several fragments through the IM than it does to have a ligase "ligate once and fall over" then that would favour ligases in the periplasm.
ReplyDelete