At one time DNA uptake was thought to be mediated by membrane-bound vesicles called transformasomes (Goodgal and Smith lab papers). These structures appear as surface 'blebs' in electron mocrographs of competent H. influenzae and H. parainfluenzae and to a lesser extent on non-competent cells. Blebs often contain DNA which is inaccessible both to externally added nucleases and to the restriction enzymes present in the cell's cytoplasm. They appear to be internalized on DNA uptake (only in H. para?), and are shed into the medium by some competence mutants and when normal cells lost competence. Purified blebs bind DNA tightly and with the same specificity as intact cells but do not internalize it (???).
The name transformasomes resulted from the interpretation that these blebs were structural adaptations for DNA uptake. However microbiologists have subsequently realized that similar blebs are often produced when gram negative bacteria are stressed (Beveridge papers), and they are no longer thought to be specialized for DNA uptake. Rather, they are thought to contain DNA uptake machinery only because this is located in thecell envelope from which the blebs form.
However, it is still possible that those parts of the envelope containing the DNA uptake machinery are particularly likely to form blebs. The proposal will describe using competence mutants to test this. Even if blebs are not enriched for DNA uptake machinery relative to the cell envelope, they lack all of the cytoplasmic proteins and this may be a valuable tool for characterizing competence-specific proteins and protein-DNA interactions. Their ability to bind DNA tightly without taking it up will be especially useful.
- Home
- Angry by Choice
- Catalogue of Organisms
- Chinleana
- Doc Madhattan
- Games with Words
- Genomics, Medicine, and Pseudoscience
- History of Geology
- Moss Plants and More
- Pleiotropy
- Plektix
- RRResearch
- Skeptic Wonder
- The Culture of Chemistry
- The Curious Wavefunction
- The Phytophactor
- The View from a Microbiologist
- Variety of Life
Field of Science
-
-
Change of address9 months ago in Variety of Life
-
Change of address9 months ago in Catalogue of Organisms
-
-
Earth Day: Pogo and our responsibility1 year ago in Doc Madhattan
-
What I Read 20241 year ago in Angry by Choice
-
I've moved to Substack. Come join me there.1 year ago in Genomics, Medicine, and Pseudoscience
-
-
-
-
Histological Evidence of Trauma in Dicynodont Tusks7 years ago in Chinleana
-
Posted: July 21, 2018 at 03:03PM7 years ago in Field Notes
-
Why doesn't all the GTA get taken up?7 years ago in RRResearch
-
-
Harnessing innate immunity to cure HIV9 years ago in Rule of 6ix
-
-
-
-
-
-
post doc job opportunity on ribosome biochemistry!11 years ago in Protein Evolution and Other Musings
-
Blogging Microbes- Communicating Microbiology to Netizens11 years ago in Memoirs of a Defective Brain
-
Re-Blog: June Was 6th Warmest Globally11 years ago in The View from a Microbiologist
-
-
-
The Lure of the Obscure? Guest Post by Frank Stahl13 years ago in Sex, Genes & Evolution
-
-
Lab Rat Moving House14 years ago in Life of a Lab Rat
-
Goodbye FoS, thanks for all the laughs14 years ago in Disease Prone
-
-
Slideshow of NASA's Stardust-NExT Mission Comet Tempel 1 Flyby15 years ago in The Large Picture Blog
-
in The Biology Files
Not your typical science blog, but an 'open science' research blog. Watch me fumbling my way towards understanding how and why bacteria take up DNA, and getting distracted by other cool questions.
Getting the kinks in
I'm finally digging in on the grant proposal planning, with the goal of having a presentable draft before Christmas. A colleague in the US has agreed to critique it for me and I want to give him lots of time, and myself lots of time to fix it up once he's done.
I think I'm going to propose to test my hypothesis that DNA forms kinks at the USS during uptake.
Because the need to invoke kinks arises because cells can take up circular DNA molecules, I wonder if we should replicate the old experiments showing that closed circular molecules are efficiently taken up. There are lots of papers where circular plasmids were transformed into competent cells, but in these there was no analysis to rule out effects of nicked or linear plasmids contaminating the preparation. But I found the best paper, and the data looks quite solid. They used a preparation of closed circular molecules that gave a ladder of bands in gels, indicative of different extents of supercoiling, and the ladder was preserved in the DNA they later reextracted from cells that had taken the plasmids up. This confirms that the circular molecules were taken up.
It's possible that the kink arises because the DNA is nicked during uptake (it's easy to bend DNA sharply at a nick), but if so the nick must be resealed without changing the degree of supercoiling. This would require a specialized nicking enzyme for which we have no evidence. However we do have evidence of a ligase in the periplasm that could reseal nicks, though no idea why this would be beneficial.
So how does DNA form kinks? This time my literature searching led me into an area I'd missed previously - the ability of very short fragments to be ligated into circles. Linear DNAs longer than 200bp are easily ligated because the molecules spontaneously bend in smooth curves that (sometimes) bring the ends together. But shorter molecules (e.g. 100bp) were thought to not do this because their smooth curves would be too short - they'd make semicircles with the ends far apart instead of complete circles.
However Jon Widom's lab showed a few years ago that 100bp fragments form circles much more efficiently than anyone had expected, and this has directed attention to how spontaneous fluctuations in DNA structure cause transient kinking. One way this can happen is by formation of one or more internal bubbles (unpaired bases), because it's the base pairing that gives DNA its stiffness. Kinks can also be promoted by DNA-binding proteins that not only undo base pairs but flip one base out of the double helix
One issue that I don't think has been resolved is the extent to which the fluctuations only happen with short fragments. The ends of any double-stranded DNA molecules are known to spontaneously 'breathe', unzipping the terminal base pairs and zipping them up again due to thermal noise. In very short fragments this could affect base pairing throughout the fragment, facilitating formation of internal bubbles and kinks.
I think I'm going to propose to test my hypothesis that DNA forms kinks at the USS during uptake.
Because the need to invoke kinks arises because cells can take up circular DNA molecules, I wonder if we should replicate the old experiments showing that closed circular molecules are efficiently taken up. There are lots of papers where circular plasmids were transformed into competent cells, but in these there was no analysis to rule out effects of nicked or linear plasmids contaminating the preparation. But I found the best paper, and the data looks quite solid. They used a preparation of closed circular molecules that gave a ladder of bands in gels, indicative of different extents of supercoiling, and the ladder was preserved in the DNA they later reextracted from cells that had taken the plasmids up. This confirms that the circular molecules were taken up.
It's possible that the kink arises because the DNA is nicked during uptake (it's easy to bend DNA sharply at a nick), but if so the nick must be resealed without changing the degree of supercoiling. This would require a specialized nicking enzyme for which we have no evidence. However we do have evidence of a ligase in the periplasm that could reseal nicks, though no idea why this would be beneficial.
So how does DNA form kinks? This time my literature searching led me into an area I'd missed previously - the ability of very short fragments to be ligated into circles. Linear DNAs longer than 200bp are easily ligated because the molecules spontaneously bend in smooth curves that (sometimes) bring the ends together. But shorter molecules (e.g. 100bp) were thought to not do this because their smooth curves would be too short - they'd make semicircles with the ends far apart instead of complete circles.
However Jon Widom's lab showed a few years ago that 100bp fragments form circles much more efficiently than anyone had expected, and this has directed attention to how spontaneous fluctuations in DNA structure cause transient kinking. One way this can happen is by formation of one or more internal bubbles (unpaired bases), because it's the base pairing that gives DNA its stiffness. Kinks can also be promoted by DNA-binding proteins that not only undo base pairs but flip one base out of the double helix
One issue that I don't think has been resolved is the extent to which the fluctuations only happen with short fragments. The ends of any double-stranded DNA molecules are known to spontaneously 'breathe', unzipping the terminal base pairs and zipping them up again due to thermal noise. In very short fragments this could affect base pairing throughout the fragment, facilitating formation of internal bubbles and kinks.
Pinning down the covariation
The Defining-the-USS paper needs one last analysis. I wrote earlier that the set of sites most likely to reveal the uptake bias is those that are neither in genes nor likely to function as transcriptional terminators. I have this set (490 sites) and I've made a logo (here).
I also did a MatrixPlot analysis to look for covariation between bases at different positions. (I've posted about using MatrixPlot to analyze the whole-genome set here.) It showed stronger interactions than those seen in the whole-genome set. But I'm left with two unresolved issues.
First, I don't know how strong the covariation is, because MatrixPlot's statistical underpinnings aren't explained (or aren't explained in a way that's accessible to my statistically untrained mind). I can take care of this by doing a control analysis, using the same number of input sequences taken from random positions in the genome - this control is on hold because the MatrixPlot server is down. (I guess really this control should use random intergenic positions - I could do that.) This is a valid control even though it doesn't use any statistics.
Second, when MatrixPlot detects covariation between two positions it doesn't tell me which combinations bases are found together more often than expected. For example, it usually reports that the base at position 18 (in the first flanking segment) is strongly correlated with the base at position 19. Both A and T are common at both of these positions, but MatrixPlot doesn't tell me whether the significant combinations are AA or AT or TA or TT.
My colleague Steve Schaeffer's linkage disequilibrium program will do that, so I'm about to email him and ask him to run the unconstrained (non-coding non-terminator) and control sets for us. But I want to get the control MatrixPlot results first, so I can more clearly explain what we think is going on.
I also did a MatrixPlot analysis to look for covariation between bases at different positions. (I've posted about using MatrixPlot to analyze the whole-genome set here.) It showed stronger interactions than those seen in the whole-genome set. But I'm left with two unresolved issues.
First, I don't know how strong the covariation is, because MatrixPlot's statistical underpinnings aren't explained (or aren't explained in a way that's accessible to my statistically untrained mind). I can take care of this by doing a control analysis, using the same number of input sequences taken from random positions in the genome - this control is on hold because the MatrixPlot server is down. (I guess really this control should use random intergenic positions - I could do that.) This is a valid control even though it doesn't use any statistics.
Second, when MatrixPlot detects covariation between two positions it doesn't tell me which combinations bases are found together more often than expected. For example, it usually reports that the base at position 18 (in the first flanking segment) is strongly correlated with the base at position 19. Both A and T are common at both of these positions, but MatrixPlot doesn't tell me whether the significant combinations are AA or AT or TA or TT.
My colleague Steve Schaeffer's linkage disequilibrium program will do that, so I'm about to email him and ask him to run the unconstrained (non-coding non-terminator) and control sets for us. But I want to get the control MatrixPlot results first, so I can more clearly explain what we think is going on.
A genuinely unbiased motif
In a previous post I showed logos from four replicate Gibbs analyses of Goodgal and Mitchell's set of plasmid insert sequences. Because the replicates didn't produce identical motif sites, the logos differed. I've now done 10 replicate runs (each testing 100 seeds) and pooled the results to create a single logo.
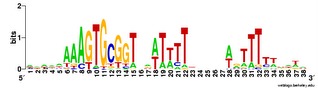 Not only did the different replicates find different numbers of sites (between 21 and 28), they settled on motifs with slightly different centers and (once) a different length. This gives me more confidence that this consensus logo represents most of the variation in sites.
Not only did the different replicates find different numbers of sites (between 21 and 28), they settled on motifs with slightly different centers and (once) a different length. This gives me more confidence that this consensus logo represents most of the variation in sites.
One remaining concern is that these searches were not unbiased, in that I started them all with a 'fragmentation mask' that specified the positions of gaps in the motif. See this post for an explanation. What would a completely unbiased search find? I suspect it will close the gaps, and ignore the consensus at positions that aren't close to the core, as this is what happens in unbiased searches of the genome. But I should try (now).
...Short break while I do this (28 short sequences means it runs very fast)...
Well I'll be hornswoggled! (Sorry, Language Log influence.) It found the reverse version of the full USS motif!
...Another slightly longer break while I do ten replicate runs...
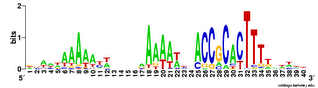 And here's two versions of the logo from the unbiased searches. The top one is what the searches find - the USS motif in reverse orientation . The lower logo is the reverse complement of this (same data, but in back-to-front order and with all the As
And here's two versions of the logo from the unbiased searches. The top one is what the searches find - the USS motif in reverse orientation . The lower logo is the reverse complement of this (same data, but in back-to-front order and with all the As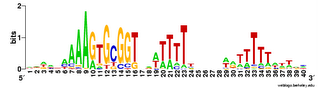  changed to Ts, Ts to As, Gs to Cs and Cs to Gs).
changed to Ts, Ts to As, Gs to Cs and Cs to Gs).
This 'unbiased' motif is actually a better match to the genome consensus than is the motif I got using the fragmentation mask. How nice!
I've been (sometimes) putting 'unbiased' in quotes, because I don't think any pattern search can be truly unbiased. The Gibbs motif sampler program I'm using for these searches has a bias towards compact motifs - there's a built-in penalty for introducing gaps.
 Not only did the different replicates find different numbers of sites (between 21 and 28), they settled on motifs with slightly different centers and (once) a different length. This gives me more confidence that this consensus logo represents most of the variation in sites.
Not only did the different replicates find different numbers of sites (between 21 and 28), they settled on motifs with slightly different centers and (once) a different length. This gives me more confidence that this consensus logo represents most of the variation in sites.One remaining concern is that these searches were not unbiased, in that I started them all with a 'fragmentation mask' that specified the positions of gaps in the motif. See this post for an explanation. What would a completely unbiased search find? I suspect it will close the gaps, and ignore the consensus at positions that aren't close to the core, as this is what happens in unbiased searches of the genome. But I should try (now).
...Short break while I do this (28 short sequences means it runs very fast)...
Well I'll be hornswoggled! (Sorry, Language Log influence.) It found the reverse version of the full USS motif!
...Another slightly longer break while I do ten replicate runs...
 And here's two versions of the logo from the unbiased searches. The top one is what the searches find - the USS motif in reverse orientation . The lower logo is the reverse complement of this (same data, but in back-to-front order and with all the As
And here's two versions of the logo from the unbiased searches. The top one is what the searches find - the USS motif in reverse orientation . The lower logo is the reverse complement of this (same data, but in back-to-front order and with all the As  changed to Ts, Ts to As, Gs to Cs and Cs to Gs).
changed to Ts, Ts to As, Gs to Cs and Cs to Gs).This 'unbiased' motif is actually a better match to the genome consensus than is the motif I got using the fragmentation mask. How nice!
I've been (sometimes) putting 'unbiased' in quotes, because I don't think any pattern search can be truly unbiased. The Gibbs motif sampler program I'm using for these searches has a bias towards compact motifs - there's a built-in penalty for introducing gaps.
The post-doc and I sat down and went over the results we have for our 'defining the USS' manuscript. In doing this I realized that my idea about the differences between intergenic and coding USSs was wrong.
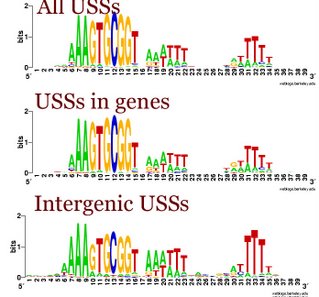 I had been thinking that the differences were due as much to the terminator function of many intergenic USS as to the coding constraints on the USSs in genes. But when we looked at the logos for the terminator and non-terminator classes of intergenic USSs, we saw that they were quite similar. Compared to the whole-genome USS logo, both showed the strong AAA at the beginning, and strong flanking motifs, especially the TTT at the end. In both the GTGCGGT of the core was also weaker. These patterns are also seen in the logo of all the intergenic USSs to the left.
I had been thinking that the differences were due as much to the terminator function of many intergenic USS as to the coding constraints on the USSs in genes. But when we looked at the logos for the terminator and non-terminator classes of intergenic USSs, we saw that they were quite similar. Compared to the whole-genome USS logo, both showed the strong AAA at the beginning, and strong flanking motifs, especially the TTT at the end. In both the GTGCGGT of the core was also weaker. These patterns are also seen in the logo of all the intergenic USSs to the left.
The USSs-in-genes logo (the sum of all 6 reading-frame specific logos I posted a week or so ago) is much more similar to the whole genome logo. This is perhaps not surprising, because most USSs are in genes.
But the similarities of the non-terminator and terminator USSs to each other means that the importance of the initial As and final Ts isn't due to selection for forming a terminator, but to the function of the sequence in DNA uptake. (On reading back over what I posted last week, I see I was pretty much thinking the same thing then. But somehow between then and this morning I got led astray.)
 I had been thinking that the differences were due as much to the terminator function of many intergenic USS as to the coding constraints on the USSs in genes. But when we looked at the logos for the terminator and non-terminator classes of intergenic USSs, we saw that they were quite similar. Compared to the whole-genome USS logo, both showed the strong AAA at the beginning, and strong flanking motifs, especially the TTT at the end. In both the GTGCGGT of the core was also weaker. These patterns are also seen in the logo of all the intergenic USSs to the left.
I had been thinking that the differences were due as much to the terminator function of many intergenic USS as to the coding constraints on the USSs in genes. But when we looked at the logos for the terminator and non-terminator classes of intergenic USSs, we saw that they were quite similar. Compared to the whole-genome USS logo, both showed the strong AAA at the beginning, and strong flanking motifs, especially the TTT at the end. In both the GTGCGGT of the core was also weaker. These patterns are also seen in the logo of all the intergenic USSs to the left.The USSs-in-genes logo (the sum of all 6 reading-frame specific logos I posted a week or so ago) is much more similar to the whole genome logo. This is perhaps not surprising, because most USSs are in genes.
But the similarities of the non-terminator and terminator USSs to each other means that the importance of the initial As and final Ts isn't due to selection for forming a terminator, but to the function of the sequence in DNA uptake. (On reading back over what I posted last week, I see I was pretty much thinking the same thing then. But somehow between then and this morning I got led astray.)
Reanalyzing old uptake data
I've been using the motif patterns identified by the Gibbs motif sampler to reanalyze old DNA uptake data. My goal is to see if the published uptake differences correlate well with motif scores that reflect match to the pattern of genomic USSs. That is, how similar is the uptake bias to the sequences that accumulate in the genome. We've independently done our own experiments to examine this correlation, but I like the notion of finding more value in old data. (A good scientist is a lazy scientist.)
So I've been using Patser (on RSAtools) to generate a motif score for each 'suitable' fragment. To be suitable, the sequence of the fragment is available (either from the original authors or from subsequent sequencing projects) and the uptake data must be quantitative. It's not enough that the original paper provided images of gels showing that some fragments were taken up better than others; I need numerical measures of uptake. In the best data (from Sol Goodgal and Marylin Mitchell) the uptake is reported as numbers of molecules taken up per cell under standard conditions. In another experiment (Danner et al 1982) uptake is presented as % relative to a standard fragment. Several other papers describe uptake results qualitatively ('strong', 'weak', 'undetectable'), and may show gels, but I can't use these.
 Here are the results. The blue and red points are data from Goodgal and Mitchell, and the green ones from Danner et al.
Here are the results. The blue and red points are data from Goodgal and Mitchell, and the green ones from Danner et al.
Goodgal tested a set of 28 plasmids for uptake, reporting the results as plasmid molecules taken up per cell. These are the blue points.; they form two clear clusters. Cells take up fewer than 30 molecules of plasmids with USS scores less than about 8, and take up between 65 and 80 molecules of plasmids with USS scores better than 9.
(Oops, I forgot to take into account the sequence of the plasmid vector the genome fragments were cloned into. This is pUC18. So I just got the pUC18 DNA sequence (Googled it), tidied it up by removing spaces and line breaks, and ran it through Patser to see if it has any USS-like sequences. This gave four surprisingly high scoring sequences - the scores are all between 10-5 and 11.5, but these sequences don't look anything like USSs to me. I'll need to think about this some more, but as all the tested plasmids were in the same vector I don't think this compromises the results.)
The red points are from the same data set. Goodgal and Mitchell purified the insert fragments from 15 of their plasmids and tested them again, this time measuring the % of the DNA that was taken up but not converting this to the number of molecules per cell. (Hmm, I wonder if I could do that myself? They say they used about 0.01µg of fragment per 0.1ml cells (about 10^8 cells, because they used 0.1ml of competent cells, and these are usually at about 10^9 cells/ml). Using Rosie's universal constant (a 1kb fragment weighs 10^-18g), and some dimensional analysis, I see that molecules taken up per cell = 1000 x %uptake/fragment size in bp.) OK, I'll go back later and make the conversion.
The green points are from Danner et al'.s analysis of synthetic USSs they constructed using the then-new ability to sequence DNA oligomers of desired sequence. Their results were reported as the amount of uptake relative to a DNA fragment that had already been shown to be taken up quite efficiently (they used this fragment as an internal control in each uptake assay they did). To conveniently fit their numbers on the graph above I multiplied each relative-uptake value by 0.5.
Does it matter that I threw in this fudge-factor of 0.5? I think not, because the numbers are all relative to an arbitrary internal standard for which I have no absolute uptake data. My main goal is to see whether better uptake correlates with a better score, and in general it does. I'm not going to try to draw any more detailed conclusions.
So I've been using Patser (on RSAtools) to generate a motif score for each 'suitable' fragment. To be suitable, the sequence of the fragment is available (either from the original authors or from subsequent sequencing projects) and the uptake data must be quantitative. It's not enough that the original paper provided images of gels showing that some fragments were taken up better than others; I need numerical measures of uptake. In the best data (from Sol Goodgal and Marylin Mitchell) the uptake is reported as numbers of molecules taken up per cell under standard conditions. In another experiment (Danner et al 1982) uptake is presented as % relative to a standard fragment. Several other papers describe uptake results qualitatively ('strong', 'weak', 'undetectable'), and may show gels, but I can't use these.
 Here are the results. The blue and red points are data from Goodgal and Mitchell, and the green ones from Danner et al.
Here are the results. The blue and red points are data from Goodgal and Mitchell, and the green ones from Danner et al.Goodgal tested a set of 28 plasmids for uptake, reporting the results as plasmid molecules taken up per cell. These are the blue points.; they form two clear clusters. Cells take up fewer than 30 molecules of plasmids with USS scores less than about 8, and take up between 65 and 80 molecules of plasmids with USS scores better than 9.
(Oops, I forgot to take into account the sequence of the plasmid vector the genome fragments were cloned into. This is pUC18. So I just got the pUC18 DNA sequence (Googled it), tidied it up by removing spaces and line breaks, and ran it through Patser to see if it has any USS-like sequences. This gave four surprisingly high scoring sequences - the scores are all between 10-5 and 11.5, but these sequences don't look anything like USSs to me. I'll need to think about this some more, but as all the tested plasmids were in the same vector I don't think this compromises the results.)
The red points are from the same data set. Goodgal and Mitchell purified the insert fragments from 15 of their plasmids and tested them again, this time measuring the % of the DNA that was taken up but not converting this to the number of molecules per cell. (Hmm, I wonder if I could do that myself? They say they used about 0.01µg of fragment per 0.1ml cells (about 10^8 cells, because they used 0.1ml of competent cells, and these are usually at about 10^9 cells/ml). Using Rosie's universal constant (a 1kb fragment weighs 10^-18g), and some dimensional analysis, I see that molecules taken up per cell = 1000 x %uptake/fragment size in bp.) OK, I'll go back later and make the conversion.
The green points are from Danner et al'.s analysis of synthetic USSs they constructed using the then-new ability to sequence DNA oligomers of desired sequence. Their results were reported as the amount of uptake relative to a DNA fragment that had already been shown to be taken up quite efficiently (they used this fragment as an internal control in each uptake assay they did). To conveniently fit their numbers on the graph above I multiplied each relative-uptake value by 0.5.
Does it matter that I threw in this fudge-factor of 0.5? I think not, because the numbers are all relative to an arbitrary internal standard for which I have no absolute uptake data. My main goal is to see whether better uptake correlates with a better score, and in general it does. I'm not going to try to draw any more detailed conclusions.
USS as terminators
I've more-or-less finished the analysis of USSs in non-coding parts of the genome. I say more-or-less because I never did get the Gibbs searches to work properly on the correct set of intergenic sequences, even after I took the advice of the Gibbs expert and replaced all the sequences less than 30nt with long strings of 'n's.
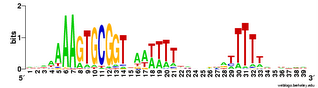
But the Gibbs searches would sometimes run correctly if I only asked them to test 1 or 2 or 3 seeds, so I got some useful data. Here are the results. This logo shows the pattern for the 490 USSs that are neither in coding sequences nor in positions where they are likely to function as transcriptional terminators. So this represents those USSs that are least functionally constrained.
For comparison, here is the logo for all the USSs Gibbs finds in the genome (2136). You can see that the initial As and final Ts are stronger (larger) in the least-constrained USSs. This also makes the USS pattern more strongly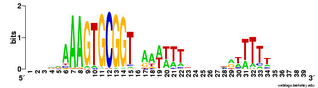 palindromic, so it is symmetric when both DNA strands are considered. To me this suggests that the DNA may kink in the middle, between positions 19 and 20, with base-pair interactions between the initial As and terminal Ts. Tomorrow morning I'm meeting with a structural biochemist who probably set me straight about this. Her main expertise is in protein structure, but at least she'll be able to point me to the best sources of information about DNA kinking.
palindromic, so it is symmetric when both DNA strands are considered. To me this suggests that the DNA may kink in the middle, between positions 19 and 20, with base-pair interactions between the initial As and terminal Ts. Tomorrow morning I'm meeting with a structural biochemist who probably set me straight about this. Her main expertise is in protein structure, but at least she'll be able to point me to the best sources of information about DNA kinking.
And here is the complementary logo, based on only the 223 USSs that are both in non-coding regions and in oppositely-oriented pairs close enough to together act as a transcription terminator. The initial As and terminal Ts are still very strong, but now we see a new ACCGCAC pattern on the right, capable of base pairing with the GTGCGGT bases on the left.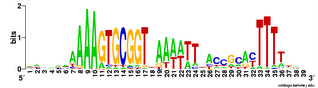 I'm going to have to think more about what this means, as I can't just say "It reflects functional constraints...". (My thinking will mostly consist of drawing sketches on the whiteboards in the hall outside my office.)
I'm going to have to think more about what this means, as I can't just say "It reflects functional constraints...". (My thinking will mostly consist of drawing sketches on the whiteboards in the hall outside my office.)

But the Gibbs searches would sometimes run correctly if I only asked them to test 1 or 2 or 3 seeds, so I got some useful data. Here are the results. This logo shows the pattern for the 490 USSs that are neither in coding sequences nor in positions where they are likely to function as transcriptional terminators. So this represents those USSs that are least functionally constrained.
For comparison, here is the logo for all the USSs Gibbs finds in the genome (2136). You can see that the initial As and final Ts are stronger (larger) in the least-constrained USSs. This also makes the USS pattern more strongly
 palindromic, so it is symmetric when both DNA strands are considered. To me this suggests that the DNA may kink in the middle, between positions 19 and 20, with base-pair interactions between the initial As and terminal Ts. Tomorrow morning I'm meeting with a structural biochemist who probably set me straight about this. Her main expertise is in protein structure, but at least she'll be able to point me to the best sources of information about DNA kinking.
palindromic, so it is symmetric when both DNA strands are considered. To me this suggests that the DNA may kink in the middle, between positions 19 and 20, with base-pair interactions between the initial As and terminal Ts. Tomorrow morning I'm meeting with a structural biochemist who probably set me straight about this. Her main expertise is in protein structure, but at least she'll be able to point me to the best sources of information about DNA kinking.And here is the complementary logo, based on only the 223 USSs that are both in non-coding regions and in oppositely-oriented pairs close enough to together act as a transcription terminator. The initial As and terminal Ts are still very strong, but now we see a new ACCGCAC pattern on the right, capable of base pairing with the GTGCGGT bases on the left.
 I'm going to have to think more about what this means, as I can't just say "It reflects functional constraints...". (My thinking will mostly consist of drawing sketches on the whiteboards in the hall outside my office.)
I'm going to have to think more about what this means, as I can't just say "It reflects functional constraints...". (My thinking will mostly consist of drawing sketches on the whiteboards in the hall outside my office.)
Oh right, the grant proposal...
Research is fun, and it's easy to get so caught up in the day-to-day problems and discoveries that I put off addressing the longer-term goals. Such as the need to prepare a compelling research proposal that will get us lots of grant money for the next 3-5 years. (Only the best proposals get money for 5 years; the rest get it for 3.)
One of the post-docs has been doing a lot of thinking about how to study the DNA uptake machinery - this will be the main focus of the proposal. Yesterday we talked about her idea of using the little membrane 'blebs' that fall off of bacterial cells as sources enriched for the proteins of the uptake machinery (a great idea). Today she asked me about genetic approaches, quoting a maxim from Stanley Maloy, a superb bacterial geneticist and one of her instructors in the Cold Spring Harbor course in Advanced Bacterial Genetics. Stanley said something to the effect of "Protein work is all very well, but there's usually a better way that uses genetics."
So we talked about a series of experiments the lab started several years ago, which aimed to use genetic complementation to identify the gene encoding the sequence-specific part of the uptake machinery. The experiments ran into various sorts of technical and personnel problems (the technician went to South America and I had other responsibilities) and never produced the results we wanted. But now I think we're going to revive them. The post-doc brought back new methods from the Cold Spring Harbor course that will make the analysis more powerful, and she has the time and skills to focus on this problem. If nothing else the plan will be a strong part of our grant proposal.
One of the post-docs has been doing a lot of thinking about how to study the DNA uptake machinery - this will be the main focus of the proposal. Yesterday we talked about her idea of using the little membrane 'blebs' that fall off of bacterial cells as sources enriched for the proteins of the uptake machinery (a great idea). Today she asked me about genetic approaches, quoting a maxim from Stanley Maloy, a superb bacterial geneticist and one of her instructors in the Cold Spring Harbor course in Advanced Bacterial Genetics. Stanley said something to the effect of "Protein work is all very well, but there's usually a better way that uses genetics."
So we talked about a series of experiments the lab started several years ago, which aimed to use genetic complementation to identify the gene encoding the sequence-specific part of the uptake machinery. The experiments ran into various sorts of technical and personnel problems (the technician went to South America and I had other responsibilities) and never produced the results we wanted. But now I think we're going to revive them. The post-doc brought back new methods from the Cold Spring Harbor course that will make the analysis more powerful, and she has the time and skills to focus on this problem. If nothing else the plan will be a strong part of our grant proposal.
New conclusions from old data
Finally I'm moving on to reanalyzing the old data. I posted about this back in August (New bottles for old wine?) but I'm only now getting back to it. The best old data is a set of 28 short DNA sequences of plasmid inserts that were preferentially taken up by competent H. influenzae cells., and the amounts of each that the cells took up. I can do two things with this data.
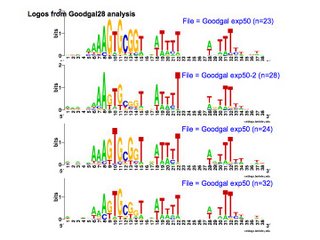 First, I can use the Gibbs motif sampler to search it for USS-like patterns. This provides one direct estimation of the bias of the uptake machinery. Here are some replicate results. All of these searches used the same dataset, but random differences in the search runs produced different datasets, which produced different logos. I haven't gone back and compared the outputs to see how many of the same sequences they found, but that will only take a few minutes to do, using Unix's 'diff' command.
First, I can use the Gibbs motif sampler to search it for USS-like patterns. This provides one direct estimation of the bias of the uptake machinery. Here are some replicate results. All of these searches used the same dataset, but random differences in the search runs produced different datasets, which produced different logos. I haven't gone back and compared the outputs to see how many of the same sequences they found, but that will only take a few minutes to do, using Unix's 'diff' command.
Second, I can find out how well the uptake of these sequences is predicted by their degree of match to the genomic USS pattern. I know that I can use a program called PatSer (for Pattern Search, I guess) on the RSA Tools site to do this. It constructs a scoring matrix and then scores sequences you give it. The matrix will be constructed from the probability matrix that the Gibbs searches produce, but I need to ask one of the graduate students to help me do this. Once I have the scores, I'll plot a graph of molecules taken up as a function of PatSer score - a strong correlation would support the hypothesis that biased uptake is responsible for the accumulation of USS in the genome.
 First, I can use the Gibbs motif sampler to search it for USS-like patterns. This provides one direct estimation of the bias of the uptake machinery. Here are some replicate results. All of these searches used the same dataset, but random differences in the search runs produced different datasets, which produced different logos. I haven't gone back and compared the outputs to see how many of the same sequences they found, but that will only take a few minutes to do, using Unix's 'diff' command.
First, I can use the Gibbs motif sampler to search it for USS-like patterns. This provides one direct estimation of the bias of the uptake machinery. Here are some replicate results. All of these searches used the same dataset, but random differences in the search runs produced different datasets, which produced different logos. I haven't gone back and compared the outputs to see how many of the same sequences they found, but that will only take a few minutes to do, using Unix's 'diff' command.Second, I can find out how well the uptake of these sequences is predicted by their degree of match to the genomic USS pattern. I know that I can use a program called PatSer (for Pattern Search, I guess) on the RSA Tools site to do this. It constructs a scoring matrix and then scores sequences you give it. The matrix will be constructed from the probability matrix that the Gibbs searches produce, but I need to ask one of the graduate students to help me do this. Once I have the scores, I'll plot a graph of molecules taken up as a function of PatSer score - a strong correlation would support the hypothesis that biased uptake is responsible for the accumulation of USS in the genome.
Palindromes Lost (when I figure out how)
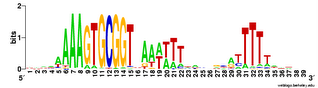 In addition to the just-posted analysis of genes in coding segments of the genome (the genes), I've analyzed the USSs in the non-coding ('intergenic') segments. Here's the logo.
In addition to the just-posted analysis of genes in coding segments of the genome (the genes), I've analyzed the USSs in the non-coding ('intergenic') segments. Here's the logo.It's a bit different than the in-gene logos in the previous post, and than the whole-genome (genes + intergenic) logo below. At first I was thinking "Great! This logo represents the unconstrained USS, because these USSs don't have to code for protein and so evolve only because of the biased DNA uptake machinery.
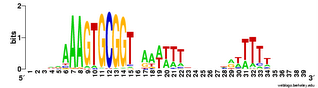
But then I realized I was forgetting about the other function that some USSs appear to have, as molecular signals marking the ends of genes. These 'stop here' signals (called transcription terminators) usually are formed by a palindromic sequence whose RNA can form a GC-rich stem-loop, followed by a short run of Ts (here's an old post that explains palindromes). USSs can do this because two USSs in opposite orientations form a potential palindrome. USSs are over-represented in non-coding regions, and they are particularly common as oppositely-oriented pairs just past the 3' ends of coding regions. RNA transcribed from one of these USSs pairs can form a GC-rich stem, and if the downstream USS is in the forward orientation this will be followed by a run of Ts. Thus these USS pairs are thought to function as transcription terminators.
Only a small subset of all USSs are in pairs that can do this, but they're all in the intergenic regions, and natural selection for the ability to terminate efficiently may constrain the effects of uptake bias. So I really ought to remove the potential terminator USSs from my intergenic dataset before I claim that it represents the unconstrained effects of uptake bias.
This is easier said than done. My initial 'clever' strategy was foiled when I realized I'd made inconsistent changes to the forward and reverse-complement versions of the intergenic sequence set. These changes didn't affect the motif search at all, but they mean that I can't use the motif-search results I have to find pairs of oppositely-oriented USSs. So I queue'd up yet more Gibbs searches, this time searching a single sequence set in both directions. These should show me where the close pairs of USSs are. Then I found a great web site that does all sorts of useful analyses for repeats. (Well, I 'found' it because a new paper from another lab, analyzing USSs in a related species, describes using this site.) The site is great, but sorting out the strands and the directions made my head spin, so I'm waiting for the Gibbs searches to finish before I decide where to apply my limited brain power.
USSs framed

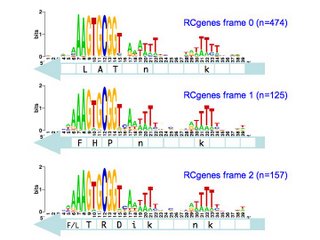
I've finished the reading frame analysis and here are the results. Each logo shows the pattern for the within-gene USSs in a specific position and orientation with respect to the reading frame of the genes they are in. The light-blue arrow below each logo shows the direction in which those USSs' genes are translated into proteins The white boxes superimposed on the arrows show the codons, with the one-letter amino acid code for those amino acids that are more- or less- strongly specified by the USS bases.
So the top left logo summarizes the 49 USSs whose positions and reverse orientation in their genes has their cores specifying the amino acids lysine (K; AAA), valine (V; GTG) and arginine (R; CGG). The consensus of the flanking regions are strong enough that they also are likely to specify particular amino acids, in this case another lysine and a phenylalanine (f; TTT). I put these last two letters in lower case to indicate that the consensus is relatively weak.
So what does this mean? We're still thinking about that. The numbers of sites (n=#, in blue) are sufficiently large that the differences between the logos are significant. I hope we will be able to correlate them to the specific coding constraints, but that's a complex analysis I'm not quite ready for yet.
Framing in progress...
Last night I did the first reading-frame analysis of USS consensuses, but only the forward three frames because I wanted to create reverse-complement sequence sets to search for the reverse three reading frames (I made the sets and ran the searches overnight). The results were interesting (yes, frame affects the consensus) but not shocking.
This morning I did the first combined analysis of all the USS in the intergenic regions. That showed more dramatic differences from the usual whole-genome consensus but I'm not sure how solid the result is, because I had only one compatible pair of forward and reverse-complement results, and both were skewed to the left of the usual motif. I have lots of results, but the forward searches almost always find a differently-centered motif than the reverse searches, so the results can't properly be compared. So I've queue'd more searches, hoping I can
This morning I thought I had plenty of results to analyze the reverse frames, until I discovered that I'd made an error in one of sequence sets (pasted in duplicates of about 100 gene sequences). So I fixed that sequence set and the searches are running again.
I also used the forward-gene and intergenic results I have to look for covariation between different USS positions in these specialized sets - nothing striking turned up.
This morning I did the first combined analysis of all the USS in the intergenic regions. That showed more dramatic differences from the usual whole-genome consensus but I'm not sure how solid the result is, because I had only one compatible pair of forward and reverse-complement results, and both were skewed to the left of the usual motif. I have lots of results, but the forward searches almost always find a differently-centered motif than the reverse searches, so the results can't properly be compared. So I've queue'd more searches, hoping I can
This morning I thought I had plenty of results to analyze the reverse frames, until I discovered that I'd made an error in one of sequence sets (pasted in duplicates of about 100 gene sequences). So I fixed that sequence set and the searches are running again.
I also used the forward-gene and intergenic results I have to look for covariation between different USS positions in these specialized sets - nothing striking turned up.
Framed?
In between thinking about the sxy manuscript (yes, it's getting closer to being done), I'm making progress on the motif analysis.
One big issue arises because about 70% of the USS are in sequences that also code for proteins. So these sequences have two different evolutionary forces acting on them. First, they must continue to code for proteins that function well - any new USS that compromise the function of the part of the proteins where they arise will be eliminated by natural selection. Second, the USS that function well in DNA uptake are preferentially replacing sequences that function less well.
 We already know that USS in coding regions are preferentially found in particular reading frames. The figure to the left shows the number of perfect USS cores in each of the 6 possible reading frames of their proteins, and also shows the amino acids the USS would specify in each reading frame (source Karlin et al. 1996 NAR 24:4263-4272). You can see that most of the 395 'forward' USS are in reading frame 3, where the USS core specifies the tripeptide SAV (amino acids serine alanine valine). A similar fraction of the 571 'reverse' USSs are in reading frame -2, where their cores specify TAL (threonine alanine leucine). In both orientations the favoured frame specifies a central versatile amino acid flanked by one hydrophilic and one hydrophobic amino acid. Thus USS whose cores encode SAL and TAV have been suggested to be common because they are better tolerated than the others.
We already know that USS in coding regions are preferentially found in particular reading frames. The figure to the left shows the number of perfect USS cores in each of the 6 possible reading frames of their proteins, and also shows the amino acids the USS would specify in each reading frame (source Karlin et al. 1996 NAR 24:4263-4272). You can see that most of the 395 'forward' USS are in reading frame 3, where the USS core specifies the tripeptide SAV (amino acids serine alanine valine). A similar fraction of the 571 'reverse' USSs are in reading frame -2, where their cores specify TAL (threonine alanine leucine). In both orientations the favoured frame specifies a central versatile amino acid flanked by one hydrophilic and one hydrophobic amino acid. Thus USS whose cores encode SAL and TAV have been suggested to be common because they are better tolerated than the others.
This analysis considered only perfect USS cores. Now I'm using the Gibbs sampler to get better datasets, and I want to find out whether being in coding regions affects the USS motif. I've finally managed to get the sampler to work on sets of genes, by splitting the gene sequence file into four short files it could handle and by specifying the expected motif structure with a strict 'prior' file.
The TIGR database provides links where I could download a file with all the coding sequences and a fine with all the non-coding sequences. Originally I thought I could only use these to get single motifs for (i) all the USS that code in the forward direction, (ii) all the USS that code in the reverse direction, and (iii) all the USS in non-coding parts of the genome. I know that it would be much more informative if I could sort out the sequences by reading frame (as in the drawing above), but I thought that would take some sophisticated bioinformatics skills.
But today I realized that it would actually be simple. The Gibbs output specifies, for each site that fits the motif, the position of its first base relative to the first base of the sequence. Because the first base is the start of the protein, I can find the reading frame simply by dividing the position number by 3. If the position of the first base of the motif is an even multiple of three (e.g. position = 93) then the motif is in frame 1. If it's not an even multiple but the remainder is 1, the motif is in frame 2. If the remainder is 2, the motif is in frame 3.
So all I need to do is paste the list of output sites into Excel (after some massaging in Word to get rid of excess spaces and separate the columns by tabs). Then I take the column of positions, use Excel's MOD function to get the remainder after division by 3, and SORT the sequences by the value in this column.
I had originally searched for USS coding in the reverse direction by searching the forward-coding sequences for the reverse motif. This isn't very satisfactory as it gives me a reverse-orientation motif, which is hard to compare with the forward one. But now I've found a web page that will take batches of sequences and convert them into their reverse complements (the sequence of the other strand), so I can analyze these for the forward motif. (Is your head spinning yet, dear reader?) Unfortunately it can only handle about 100 sequences at once, so I have to do the whole set (about 1730) in parts.
So that's what I'm about to do. I think I have all the Gibbs output files I need for this. The outputs aren't always centered on the same USS position so they're not all equally useful, but I think I have complete sets with the same center. And I have a few more queue'd up just in case.
One big issue arises because about 70% of the USS are in sequences that also code for proteins. So these sequences have two different evolutionary forces acting on them. First, they must continue to code for proteins that function well - any new USS that compromise the function of the part of the proteins where they arise will be eliminated by natural selection. Second, the USS that function well in DNA uptake are preferentially replacing sequences that function less well.
 We already know that USS in coding regions are preferentially found in particular reading frames. The figure to the left shows the number of perfect USS cores in each of the 6 possible reading frames of their proteins, and also shows the amino acids the USS would specify in each reading frame (source Karlin et al. 1996 NAR 24:4263-4272). You can see that most of the 395 'forward' USS are in reading frame 3, where the USS core specifies the tripeptide SAV (amino acids serine alanine valine). A similar fraction of the 571 'reverse' USSs are in reading frame -2, where their cores specify TAL (threonine alanine leucine). In both orientations the favoured frame specifies a central versatile amino acid flanked by one hydrophilic and one hydrophobic amino acid. Thus USS whose cores encode SAL and TAV have been suggested to be common because they are better tolerated than the others.
We already know that USS in coding regions are preferentially found in particular reading frames. The figure to the left shows the number of perfect USS cores in each of the 6 possible reading frames of their proteins, and also shows the amino acids the USS would specify in each reading frame (source Karlin et al. 1996 NAR 24:4263-4272). You can see that most of the 395 'forward' USS are in reading frame 3, where the USS core specifies the tripeptide SAV (amino acids serine alanine valine). A similar fraction of the 571 'reverse' USSs are in reading frame -2, where their cores specify TAL (threonine alanine leucine). In both orientations the favoured frame specifies a central versatile amino acid flanked by one hydrophilic and one hydrophobic amino acid. Thus USS whose cores encode SAL and TAV have been suggested to be common because they are better tolerated than the others.This analysis considered only perfect USS cores. Now I'm using the Gibbs sampler to get better datasets, and I want to find out whether being in coding regions affects the USS motif. I've finally managed to get the sampler to work on sets of genes, by splitting the gene sequence file into four short files it could handle and by specifying the expected motif structure with a strict 'prior' file.
The TIGR database provides links where I could download a file with all the coding sequences and a fine with all the non-coding sequences. Originally I thought I could only use these to get single motifs for (i) all the USS that code in the forward direction, (ii) all the USS that code in the reverse direction, and (iii) all the USS in non-coding parts of the genome. I know that it would be much more informative if I could sort out the sequences by reading frame (as in the drawing above), but I thought that would take some sophisticated bioinformatics skills.
But today I realized that it would actually be simple. The Gibbs output specifies, for each site that fits the motif, the position of its first base relative to the first base of the sequence. Because the first base is the start of the protein, I can find the reading frame simply by dividing the position number by 3. If the position of the first base of the motif is an even multiple of three (e.g. position = 93) then the motif is in frame 1. If it's not an even multiple but the remainder is 1, the motif is in frame 2. If the remainder is 2, the motif is in frame 3.
So all I need to do is paste the list of output sites into Excel (after some massaging in Word to get rid of excess spaces and separate the columns by tabs). Then I take the column of positions, use Excel's MOD function to get the remainder after division by 3, and SORT the sequences by the value in this column.
I had originally searched for USS coding in the reverse direction by searching the forward-coding sequences for the reverse motif. This isn't very satisfactory as it gives me a reverse-orientation motif, which is hard to compare with the forward one. But now I've found a web page that will take batches of sequences and convert them into their reverse complements (the sequence of the other strand), so I can analyze these for the forward motif. (Is your head spinning yet, dear reader?) Unfortunately it can only handle about 100 sequences at once, so I have to do the whole set (about 1730) in parts.
So that's what I'm about to do. I think I have all the Gibbs output files I need for this. The outputs aren't always centered on the same USS position so they're not all equally useful, but I think I have complete sets with the same center. And I have a few more queue'd up just in case.
I've spent so much time fiddling with the MatrixPlot settings to get the best visualization of the correlation analysis that Matrix Plot won't let me submit any more jobs. (Who knew the site had a limit of 50 jobs per 24 hr period?)
I've done the final analysis with a set of 3466 sequences, each 39bp) and each containing at least a rough match to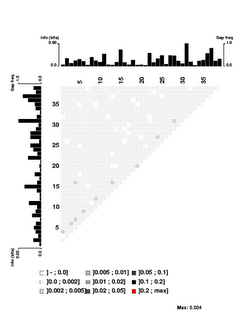 the USS motif. These were obtained by motif searches told that 3000 sites were expect on each strand; 1650 and 1816 were found. 1454 of these contain perfect matches to the 9bp core consensus, 512 have one-off matches, 557 have two-off matches, and 943 have core sequences that match the consensus at no more than 6 places. I think having this many mismatched sequences gives the analysis the power to detect correlations even between the highly-conserved core positions.
the USS motif. These were obtained by motif searches told that 3000 sites were expect on each strand; 1650 and 1816 were found. 1454 of these contain perfect matches to the 9bp core consensus, 512 have one-off matches, 557 have two-off matches, and 943 have core sequences that match the consensus at no more than 6 places. I think having this many mismatched sequences gives the analysis the power to detect correlations even between the highly-conserved core positions.
First, look at the control figure to the left. This shows analysis of 3500 random sequences, each 39bp long, taken from random segments of both strands of the H. influenzae genome. The bar charts at the top and left can be ignored - they show the 'information' at each position, but the scale for these bars only goes from 0.0 to 0.00 (weird, I know. I guess '0.0' represents zero, and '0.00' represents less than 0.01),
It's a bit surprising (to me) that the few scores higher than 0.002 are mostly found between positions separated by 3 (positions 1 and 4, positions 3 and 6, positions 4 and 7, positions 9 and 12, etc.). I suspect this has something to do with the way coding for proteins constrains the genome, but it's not something I'm going to follow up.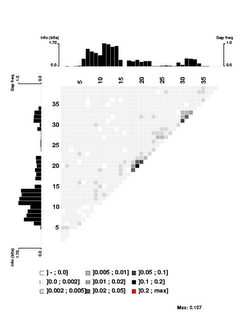
Here's the 'experimental' image. It shows significant correlations only between close-neighbour positions, and only between neighbours within each of the two flanking conserved AT segments. I suspect that even these 'significant' correlations are quite weak (the highest correlation score is only 0.107), but I don't understand the analysis well enough to be sure. The documentation is very brief; I may need to send someone an email asking for clarification.
(Here's a logo as a reminder of the motif.)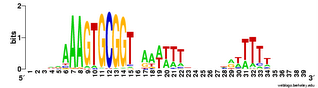
I've done the final analysis with a set of 3466 sequences, each 39bp) and each containing at least a rough match to
 the USS motif. These were obtained by motif searches told that 3000 sites were expect on each strand; 1650 and 1816 were found. 1454 of these contain perfect matches to the 9bp core consensus, 512 have one-off matches, 557 have two-off matches, and 943 have core sequences that match the consensus at no more than 6 places. I think having this many mismatched sequences gives the analysis the power to detect correlations even between the highly-conserved core positions.
the USS motif. These were obtained by motif searches told that 3000 sites were expect on each strand; 1650 and 1816 were found. 1454 of these contain perfect matches to the 9bp core consensus, 512 have one-off matches, 557 have two-off matches, and 943 have core sequences that match the consensus at no more than 6 places. I think having this many mismatched sequences gives the analysis the power to detect correlations even between the highly-conserved core positions.First, look at the control figure to the left. This shows analysis of 3500 random sequences, each 39bp long, taken from random segments of both strands of the H. influenzae genome. The bar charts at the top and left can be ignored - they show the 'information' at each position, but the scale for these bars only goes from 0.0 to 0.00 (weird, I know. I guess '0.0' represents zero, and '0.00' represents less than 0.01),
It's a bit surprising (to me) that the few scores higher than 0.002 are mostly found between positions separated by 3 (positions 1 and 4, positions 3 and 6, positions 4 and 7, positions 9 and 12, etc.). I suspect this has something to do with the way coding for proteins constrains the genome, but it's not something I'm going to follow up.

Here's the 'experimental' image. It shows significant correlations only between close-neighbour positions, and only between neighbours within each of the two flanking conserved AT segments. I suspect that even these 'significant' correlations are quite weak (the highest correlation score is only 0.107), but I don't understand the analysis well enough to be sure. The documentation is very brief; I may need to send someone an email asking for clarification.
(Here's a logo as a reminder of the motif.)

Gibbs motif search progress continues unabated
The sxy manuscript has been on hold, partly because one of the two grad students involved in it was in the far north. But he's back, and the manuscript is close enough to being finished that I'm hopeful it will be done soon ('soon' being an elastic term here). So I need to switch my attention to it and away from the motif searches for the USS-defined manuscript that have been consuming my brain power lately.
But before I stop I'm seeing how much I can get finished. I ran and analyzed the leading-strand and lagging-strand searches - their motifs are indeed identical to the composite one I posted.
The more-stringent and less-stringent searches gave the results I expected (fewer and more sites with the motif, with stronger and poorer mean scores, respectively). I used the run that gave the most sites and the worst mean scores to do a correlation analysis. (Having more sites that are imperfectly matched to the consensus increases the power of this analysis to detect weak interactions.)
The goal of the correlation analysis is to find out whether the bases at different positions of the USS interact. For example, the most common bases at position 17 are A and G, and the most common bases at position 21 are T and A. If we find that the individual USSs that have A at position 17 usually have T at position 21, and those that have G at position 17 usually have A at position 21, we would conclude that the bases at these positions interact during DNA uptake. Said another way, we'd conclude that USSs with a G at position 17 function better if they have a A at position 21.
Results: MatrixPlot found only weak correlations between only a few adjacent positions in two clusters. A colleague has kindly used software he wrote to also analyze a preliminary data set for us; I'm going to ask him if he can test the big set. Before doing this, I realized that I only have half the data, as I only did the low stringency searches on the forward strand. So I've queue'd up more searches, with the same and even lower stringency, on both forward and reverse-complement strands.
And, finally, some of the gene searches are working, thanks to fine-tuning advice from the helpful Gibbs expert. These runs are searching the sequences of only the parts of the genome that code for proteins, to see if the direction of coding affects the motif. I had to split the gene set into four parts, and two of these managed to find their motifs. So I've queue'd up more replicates, using more seeds, and also runs looking for the reverse-strand motif.
And last night I read over the Introduction and improved parts of it, though it still needs more work.
But before I stop I'm seeing how much I can get finished. I ran and analyzed the leading-strand and lagging-strand searches - their motifs are indeed identical to the composite one I posted.
The more-stringent and less-stringent searches gave the results I expected (fewer and more sites with the motif, with stronger and poorer mean scores, respectively). I used the run that gave the most sites and the worst mean scores to do a correlation analysis. (Having more sites that are imperfectly matched to the consensus increases the power of this analysis to detect weak interactions.)
The goal of the correlation analysis is to find out whether the bases at different positions of the USS interact. For example, the most common bases at position 17 are A and G, and the most common bases at position 21 are T and A. If we find that the individual USSs that have A at position 17 usually have T at position 21, and those that have G at position 17 usually have A at position 21, we would conclude that the bases at these positions interact during DNA uptake. Said another way, we'd conclude that USSs with a G at position 17 function better if they have a A at position 21.
Results: MatrixPlot found only weak correlations between only a few adjacent positions in two clusters. A colleague has kindly used software he wrote to also analyze a preliminary data set for us; I'm going to ask him if he can test the big set. Before doing this, I realized that I only have half the data, as I only did the low stringency searches on the forward strand. So I've queue'd up more searches, with the same and even lower stringency, on both forward and reverse-complement strands.
And, finally, some of the gene searches are working, thanks to fine-tuning advice from the helpful Gibbs expert. These runs are searching the sequences of only the parts of the genome that code for proteins, to see if the direction of coding affects the motif. I had to split the gene set into four parts, and two of these managed to find their motifs. So I've queue'd up more replicates, using more seeds, and also runs looking for the reverse-strand motif.
And last night I read over the Introduction and improved parts of it, though it still needs more work.
Progress continues
The computer cluster guys were great. They took extra pains to make sure we understood the big issues, they were really helpful in suggesting ways to optimize our runs, and they even gave us a tour of the WestGrid system.
 I've cataloged the reverse-complement runs (same basic results as the forward runs), and made this composite logo based on all the USSs in the genome (i.e. both strands). It doesn't look any different than a composite logo I posted about 6 weeks ago; the difference here is that now I know I've done the right analysis.
I've cataloged the reverse-complement runs (same basic results as the forward runs), and made this composite logo based on all the USSs in the genome (i.e. both strands). It doesn't look any different than a composite logo I posted about 6 weeks ago; the difference here is that now I know I've done the right analysis.
Now on to analyzing the leading-strand and lagging-strand searches. I already did a quick-and-dirty version of that too, but again now I'll have done it well.
 I've cataloged the reverse-complement runs (same basic results as the forward runs), and made this composite logo based on all the USSs in the genome (i.e. both strands). It doesn't look any different than a composite logo I posted about 6 weeks ago; the difference here is that now I know I've done the right analysis.
I've cataloged the reverse-complement runs (same basic results as the forward runs), and made this composite logo based on all the USSs in the genome (i.e. both strands). It doesn't look any different than a composite logo I posted about 6 weeks ago; the difference here is that now I know I've done the right analysis.Now on to analyzing the leading-strand and lagging-strand searches. I already did a quick-and-dirty version of that too, but again now I'll have done it well.
How different are the replicates?
I haven't solved the random-number problem (runs submitted together get the identical seed) because it's not my code (I only have the compiled file). But I'm easily circumventing it by not submitting replicate runs at the same time. This afternoon the post-doc and I meet with the local computer-cluster experts, who will no doubt be stupefied by our ignorance of things Unix. If we're lucky they'll speak English as well as Unix; if not I'll just be brazen in insisting that we understand their explanations. (It's in their own interest; otherwise they risk that we'll unintentionally do something that compromises their system.)
I'm making lots of progress with the motif search results this system has given me so far. I have at least four replicate runs of each of four kinds of searches: forward DNA strand, reverse-complement DNA strand, leading strand, lagging strand (leading and lagging refer to the direction DNA polymerase moves on the strand during DNA replication). I've analyzed only the five forward strand files so far, because I want to be clear about what analyses are worth doing before going on to the other strands' files.
I've analyzed them for:
If I'm right, the expect-2000 runs will give us datasets with lots more sites, and those sites will be mainly poorer matches to the pattern than the sites we have now. This will be useful because it may give us more power when we look for correlations between the bases at different positions. (I may have posted about this; I know the post-doc has.)
I'm making lots of progress with the motif search results this system has given me so far. I have at least four replicate runs of each of four kinds of searches: forward DNA strand, reverse-complement DNA strand, leading strand, lagging strand (leading and lagging refer to the direction DNA polymerase moves on the strand during DNA replication). I've analyzed only the five forward strand files so far, because I want to be clear about what analyses are worth doing before going on to the other strands' files.
I've analyzed them for:
- final log MAP score (all between 5096 and 5109).
- number of sites found (all between 1053 and 1058).
- how many of these sites contain perfect, singly-mismatched (one-off) or doubly mismatches (two-off) USS cores (usually 719, 205 and 105 respectively).
- the mean score of the ~1055 sites each search found (between 0.92 and 0.93). Most sites have scores very close to 1.0, but some are much lower and drive the mean down. (Hmm, I should check the median as well as the mean.)
- the number of distinct sites between each pair of searches (1 vs 2, 1 vs 3 etc.). This ranged between 5 and 17, meaning that more than 98% of the sites each search found were also found by the replicate searches. The sites found in some sites and not others were usually ones with very low scores, usually about 0.5. A few were closely spaced sites with strong scores - I suspect the program can't handle overlapping sites.
If I'm right, the expect-2000 runs will give us datasets with lots more sites, and those sites will be mainly poorer matches to the pattern than the sites we have now. This will be useful because it may give us more power when we look for correlations between the bases at different positions. (I may have posted about this; I know the post-doc has.)
Subscribe to:
Comments (Atom)















